Clear Sky Science · en
Application of machine learning and genomics for orphan crop improvement
Hidden Crops with Big Potential
Across Africa, Asia, and Latin America, millions of people rely on so‑called “orphan crops” such as sorghum, teff, cassava, and groundnut. These plants rarely make headlines, yet they often withstand heat, drought, pests, and poor soils better than global staples like wheat or rice. This review article explores how two powerful tools—genomics and machine learning—can unlock the potential of these overlooked crops, boosting local food security while also providing valuable genes that could strengthen major crops worldwide.
Why Overlooked Crops Matter
Orphan crops are sometimes called “neglected” or “underutilized” because they have received far less scientific and commercial attention than big export crops. Still, they are nutritional mainstays for many communities and are often grown in harsh, marginal environments where other crops fail. Unlike wheat or rice, most orphan crops missed out on the Green Revolution’s breeding advances and modern tools such as marker‑assisted breeding and genome editing. Genomic projects like the African Orphan Crops Consortium are beginning to sequence and catalog their DNA, but turning raw genetic data into practical improvements remains a major challenge.
Teaching Computers to Read Plants
Machine learning—computer methods that learn patterns from large datasets—is already transforming breeding in major crops. By combining genome sequences, weather and soil records, sensor readings, and images from drones or smartphones, algorithms can predict complex traits like yield, disease resistance, or grain quality. Different model types, from decision trees to deep neural networks, excel in different settings. Sometimes traditional statistical tools still match or beat deep learning, but overall, blending multiple data sources and models tends to give breeders more accurate and consistent predictions than any single approach alone.
Making the Most of Scarce Data
For orphan crops, the key obstacle is not computer power but data scarcity. Only a handful of public genomic and image collections exist, and few are large enough for conventional machine learning pipelines. Even so, first demonstrations are promising. In sorghum, for example, deep learning models used simple photographs of grain to predict protein and antioxidant levels with high accuracy, offering a cheaper alternative to lab tests. In another case, near‑infrared light measurements and deep learning were used to estimate nutritional traits in the herb Perilla. The review argues that building shared databases of genomes, images, and chemical profiles for orphan crops would quickly multiply the impact of such tools.
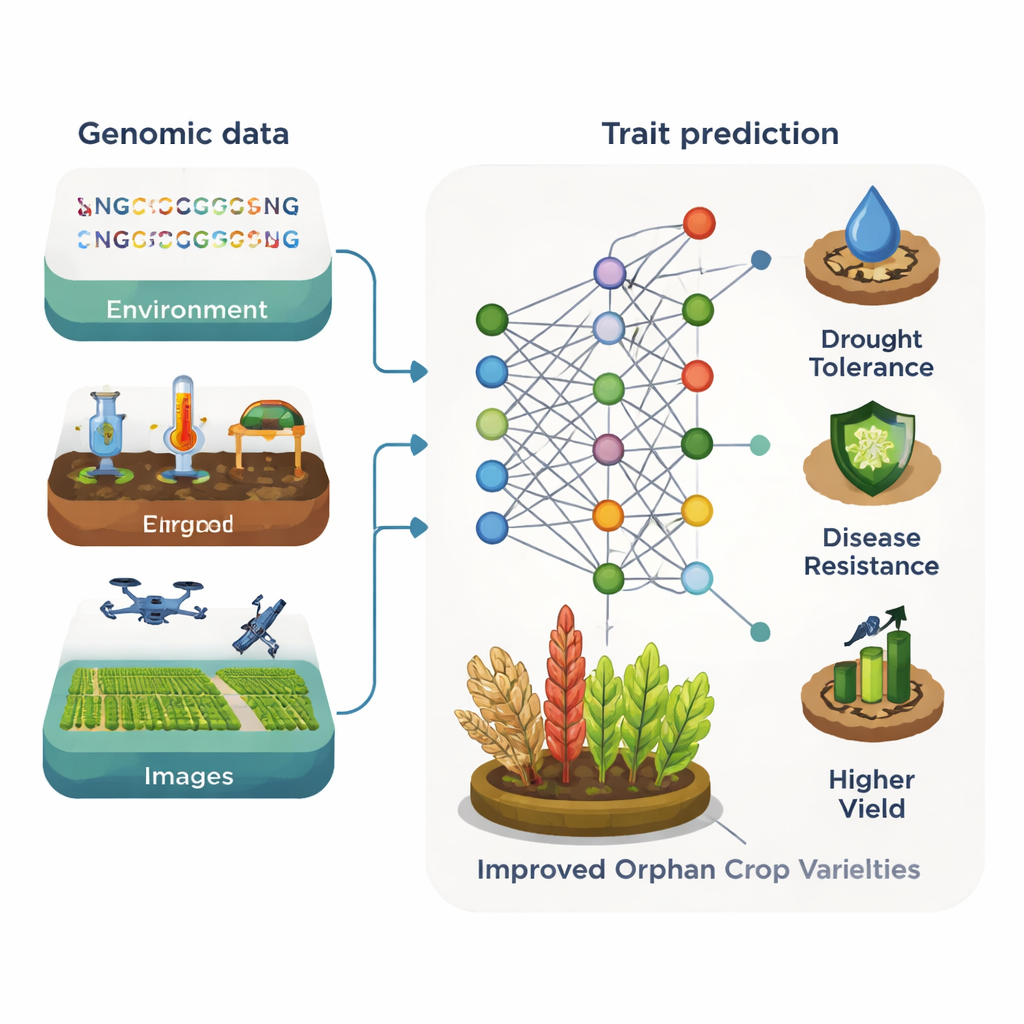
Borrowing Knowledge from Bigger Crops
A central idea of the article is “knowledge transfer” between species. Many orphan crops are close relatives of major crops, sharing large stretches of DNA and similar genes. Machine learning models can exploit this relatedness. Tools first trained on well‑studied plants like Arabidopsis or maize can help pinpoint genes for traits such as plant height, seed quality, or stress tolerance in a lesser‑known cousin. Large language models originally developed for human or plant genomes can also treat DNA as a kind of text, learning patterns that mark regulatory regions or important genes. Once trained on rich datasets, these models can be fine‑tuned on limited orphan‑crop data to predict gene function, highlight targets for genome editing, and guide more efficient breeding.
From Algorithms to Fields and Farmers
The authors stress that technology alone will not transform orphan crops. Progress depends on investment in local scientists, partnerships with smallholder farmers, and policies that ensure communities benefit from new varieties. Citizen‑science approaches, where farmers test varieties directly on their own land, can generate valuable data for machine learning while aligning research with local needs and tastes. Because funding is limited, the article recommends a balanced strategy: combine low‑cost traditional breeding and agronomy with carefully targeted genomic and machine learning projects, and share tools and data between countries and between orphan and major crops.
What This Means for Our Food Future
In plain terms, the article concludes that smarter computers plus better genetic information can help turn today’s “forgotten” crops into tomorrow’s climate‑ready staples. By learning from big crops and applying those lessons to smaller ones—and then feeding discoveries back in the other direction—machine learning and genomics can speed up the search for hardy, nutritious varieties. If supported by thoughtful policy and genuine collaboration with farming communities, this approach could improve diets, strengthen resilience to climate change, and broaden the world’s agricultural toolkit beyond a narrow set of staple crops.
Citation: MacNish, T.R., Danilevicz, M.F., Bayer, P.E. et al. Application of machine learning and genomics for orphan crop improvement. Nat Commun 16, 982 (2025). https://doi.org/10.1038/s41467-025-56330-x
Keywords: orphan crops, machine learning, genomics, crop breeding, food security