Clear Sky Science · en
Categorization of 34 computational methods to detect spatially variable genes from spatially resolved transcriptomics data
Why location matters for our genes
Our bodies are built from cells that not only differ in what they do, but also in where they sit inside tissues and organs. New "spatial transcriptomics" technologies can now read out which genes are active while keeping track of each cell’s address on a tissue map. This review article explains how scientists detect genes whose activity changes from place to place—so‑called spatially variable genes—and why agreeing on how to find them is vital for understanding cancer, brain function, and many other diseases.
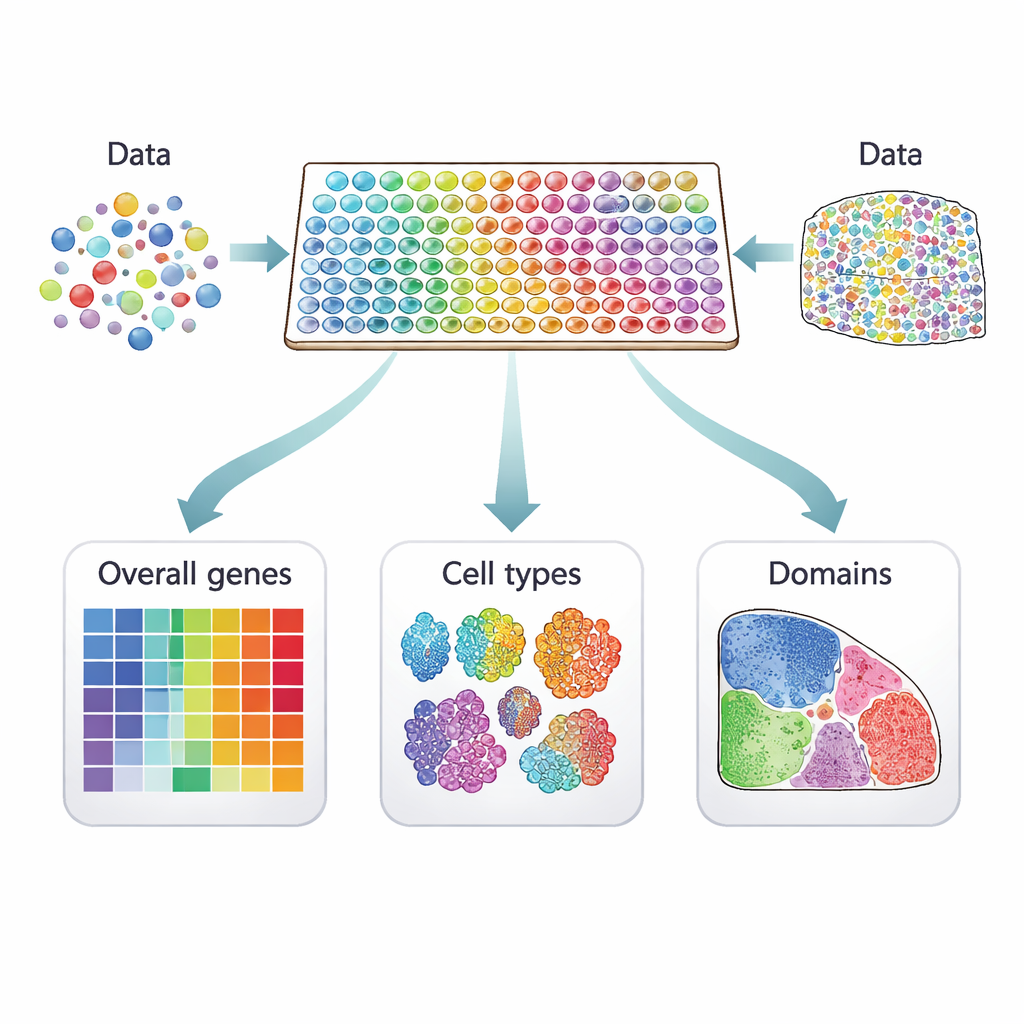
From scattered cells to living maps
Traditional single‑cell studies measure gene activity in thousands of individual cells but lose information about where those cells came from. Spatial transcriptomics fills this gap by measuring gene activity directly on thin slices of tissue. Each measurement is tied to a "spot" on the slice, which may contain one cell or several, depending on the technology. Imaging‑based methods pinpoint a few hundred chosen genes at very high spatial detail, while sequencing‑based platforms capture almost all genes at lower resolution. Together, these approaches turn a tissue section into a colorful map of gene activity that can reveal hidden structures, such as layers in the brain or regions inside a tumor.
Three kinds of location‑sensitive genes
Many computer methods have been proposed to pick out genes that show meaningful patterns across these tissue maps, but they do not all look for the same thing. The authors sort 34 current methods into three clear categories. The first group looks for "overall" spatially variable genes, whose activity changes across the tissue in any non‑random way—perhaps forming stripes, clusters, or gradients. The second group focuses on "cell‑type‑specific" spatial genes that vary within a single kind of cell, helping to distinguish, for example, subtypes of neurons or different states of immune cells. The third group searches for "domain‑marker" genes that are strongly turned on in particular regions or layers, making them useful labels for those tissue zones.
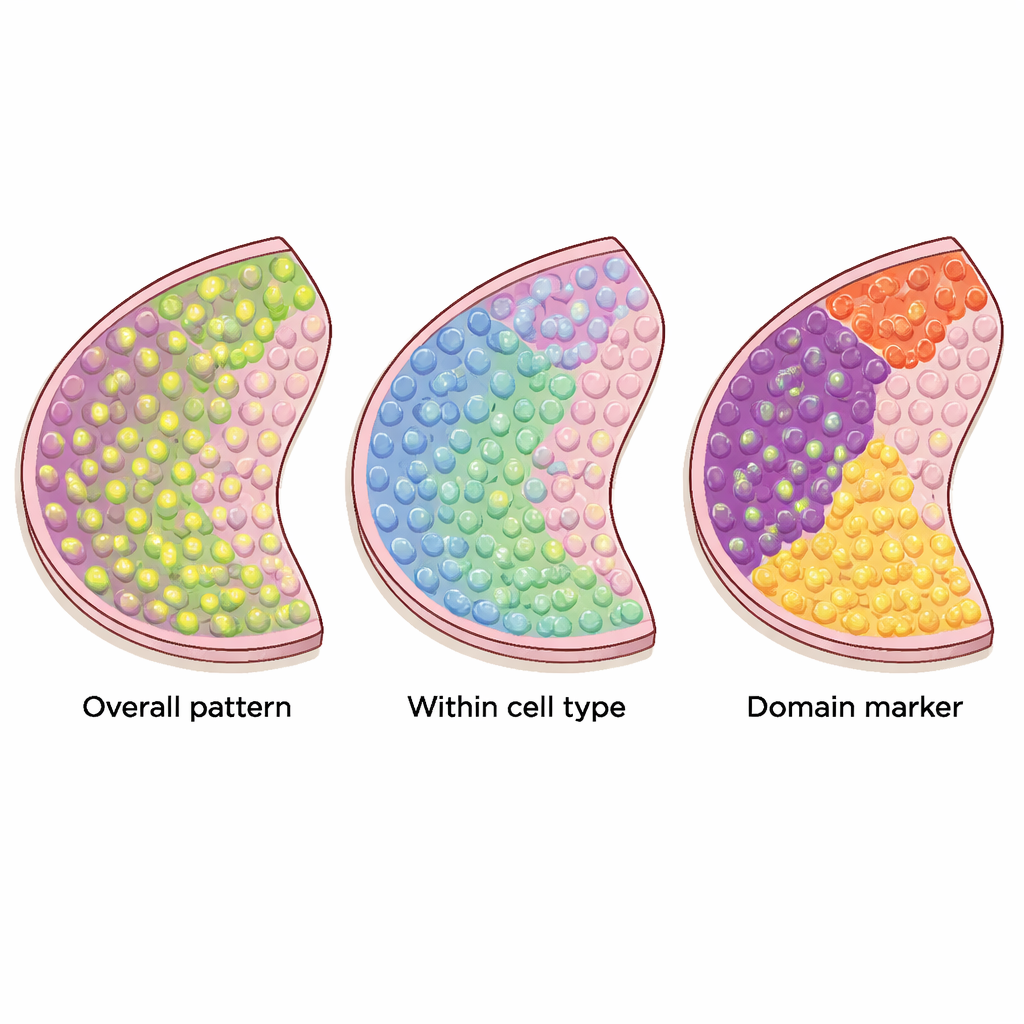
Different tools for different patterns
The review explains how these methods work under the hood. Some treat the tissue slice as points in regular two‑dimensional space and use mathematical "kernels" to look for patterns like spots or waves. Others first link nearby locations into a network, or graph, and then ask whether high expression tends to cluster along the links of that graph. Certain tools rely on formal statistical tests with well‑defined error rates, while others mainly rank genes by how strongly their patterns stand out. Methods that target very specific patterns can be powerful when the data match their expectations, but they may miss more irregular or complex shapes, such as those seen in many cancers. There is also a trade‑off between flexibility and speed: some approaches scale to hundreds of thousands of spots, while others struggle with very large datasets.
What these genes can reveal
Once spatially variable genes are identified, they become the raw material for deeper biological discoveries. Overall spatial genes are often used as a first filter to reduce the number of genes before grouping spots into "spatial domains"—regions whose cells share similar expression profiles. These domains can line up with known tissue structures, suggest new sub‑regions, or highlight distinct cell neighborhoods such as invasive fronts in tumors. Domain‑marker genes then help explain what makes each region unique and can be reused to label similar structures in other samples. Meanwhile, cell‑type‑specific spatial genes promise a finer view of how particular cell types change across a tissue, which may illuminate tumor–immune interactions or specialized circuits in the brain.
Challenges and the road ahead
The authors emphasize that no single method is best for every task, and that comparing tools fairly requires careful thought about what kind of spatial gene each method is actually designed to find. They call for better benchmarks using realistic datasets, clearer statistical standards to avoid false discoveries, and new approaches that respect differences between technologies and tissue types. For non‑experts, the key message is that spatially variable genes turn flat gene lists into living maps, linking molecular activity to tissue structure. Robust ways to detect and interpret these genes will be central to turning spatial transcriptomics into practical insights about development, brain function, and disease.
Citation: Yan, G., Hua, S.H. & Li, J.J. Categorization of 34 computational methods to detect spatially variable genes from spatially resolved transcriptomics data. Nat Commun 16, 1141 (2025). https://doi.org/10.1038/s41467-025-56080-w
Keywords: spatial transcriptomics, spatially variable genes, gene expression patterns, tissue microenvironments, computational genomics