Clear Sky Science · en
Rethinking genomics of facioscapulohumeral muscular dystrophy in the telomere-to-telomere era: pitfalls in the hidden landscape of D4Z4 repeats
Why this muscle disease story matters
Facioscapulohumeral muscular dystrophy (FSHD) is one of the most common inherited muscle-wasting diseases, often weakening the face and shoulder muscles of otherwise healthy young adults. For years, scientists have focused on a single stretch of DNA near the tip of chromosome 4 as the culprit. This new study shows that the picture is far more complicated: similar DNA repeats are scattered across the genome, and many standard lab tests cannot tell them apart. Understanding this hidden landscape is essential for accurate diagnosis, better research, and ultimately safer therapies.
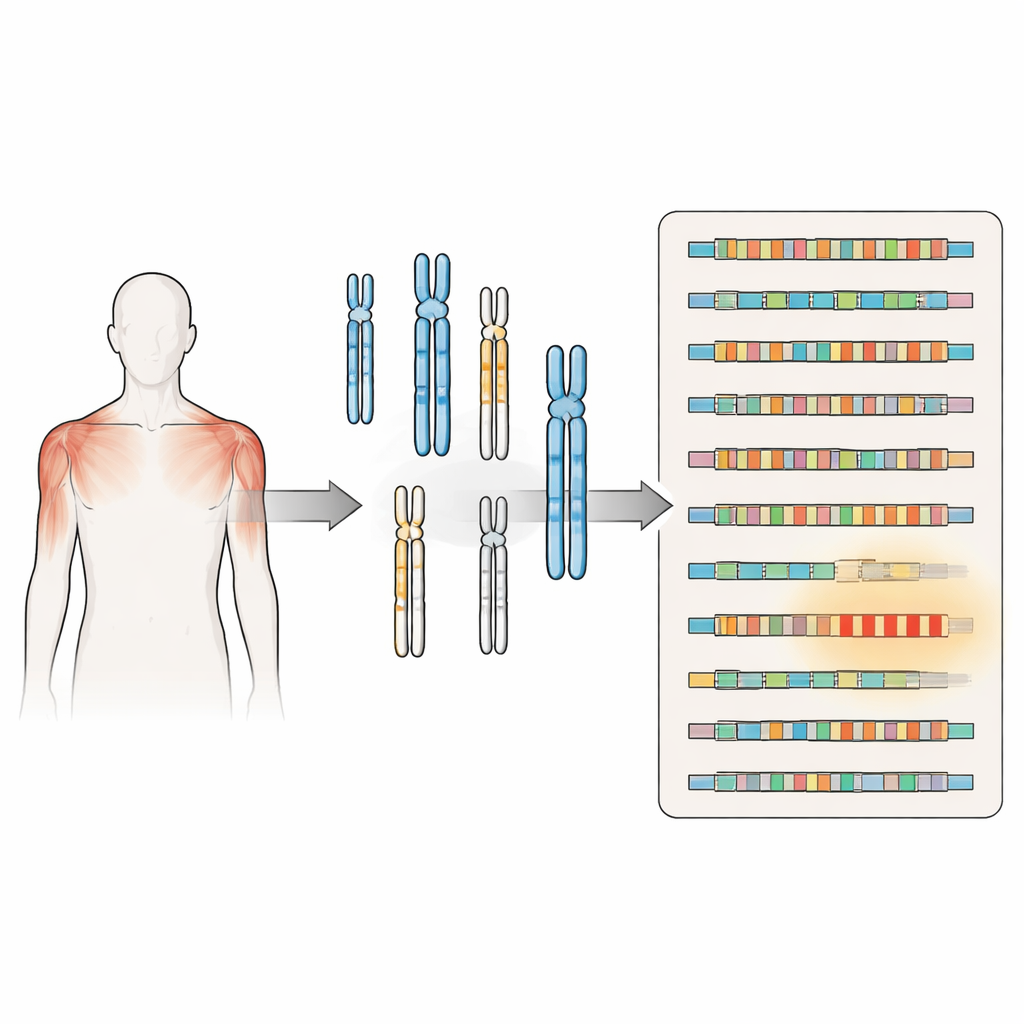
A closer look at a puzzling muscle disorder
FSHD has long been linked to the shortening of a repeated DNA sequence called D4Z4 near the end of chromosome 4. When this repeat region shrinks below a certain size and sits on a particular genetic background, it allows production of a protein called DUX4, which can damage muscle cells. A second form of the disease arises when genes that normally keep this region tightly packed and silent are mutated, again permitting DUX4 to appear. Both routes converge on the same problem: a gene that should be mostly off in adult muscle becomes sporadically active, contributing to muscle weakness and wasting.
Hidden repeats across the genome
Earlier human genome maps were incomplete, especially in regions rich in repetitive DNA at chromosome ends and near centromeres. Using the new telomere-to-telomere human genome assembly, which fills in these gaps, the authors re-scanned the entire genome using the D4Z4 sequence from chromosome 4 as a search template. They discovered clusters and isolated copies of D4Z4-like repeats on at least ten additional chromosomes. Some of these repeats are structurally complete and sit next to signals that could stabilize DUX4-like RNAs, suggesting that they might produce related proteins or RNA molecules under the right conditions.
When lab tests see more than one place at once
FSHD research and diagnosis often rely on tests that amplify specific DNA or RNA fragments with short primer sequences, a strategy known as PCR. These primers were originally designed under the assumption that D4Z4 repeats existed only on chromosomes 4 and 10. By combining computer predictions with experiments on cells that carry single human chromosomes, the authors show that many widely used primer sets bind not just to the disease-linked region on chromosome 4, but also to look‑alike repeats scattered around the genome. Primers aimed at the main DUX4 exon or at neighboring long non-coding RNAs frequently amplify products from several chromosomes at once, making it impossible to know where a given signal truly originated.
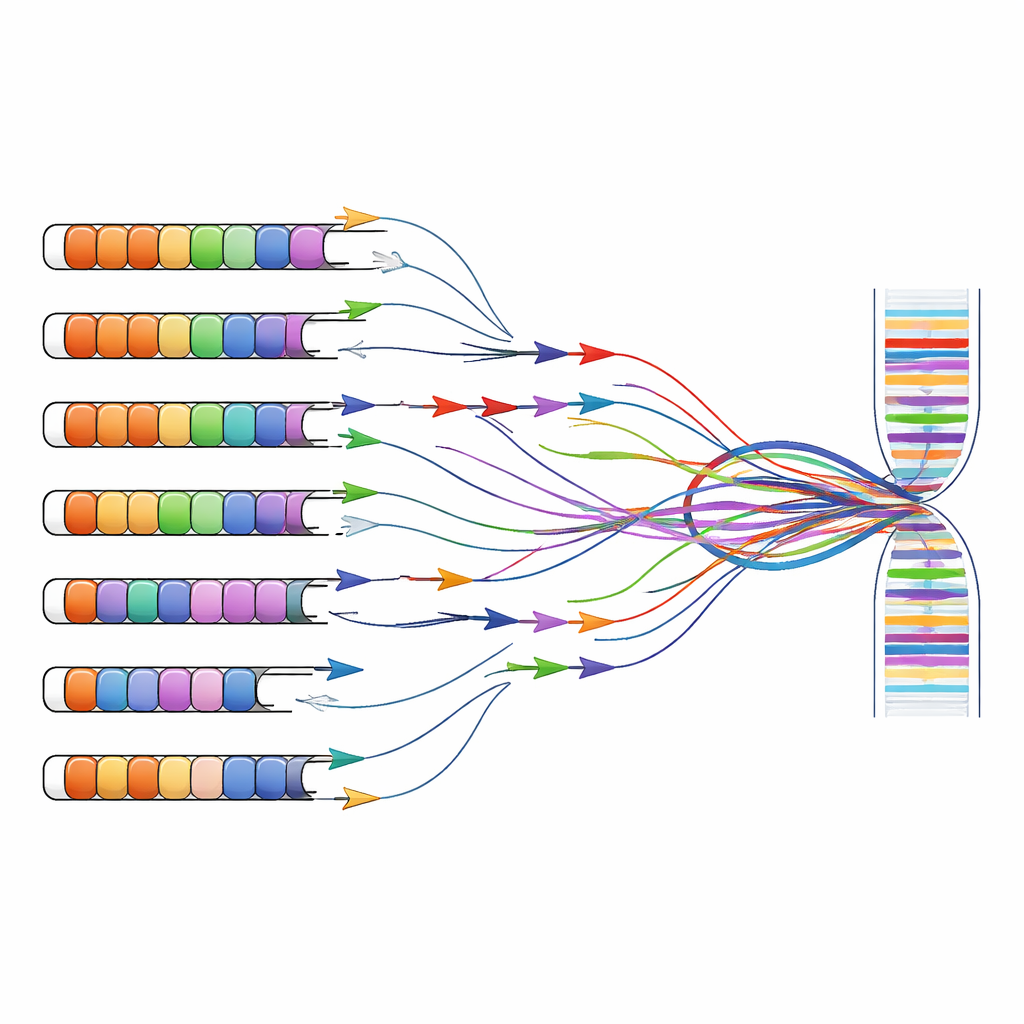
Rethinking diagnostic and research tools
This multi-location problem extends beyond basic research into the clinic. Newer tests that measure DNA methylation (a chemical mark associated with gene silencing) or use short-read sequencing often lump signals from many near-identical repeats together. As a result, measurements of “D4Z4 methylation” or “DUX4 transcripts” in public datasets may actually reflect a mixture of contributions from chromosome 4, chromosome 10, and other sites. The authors argue that to correctly link genetic changes to symptoms, scientists must now adopt repeat-aware methods that distinguish individual loci—for example, long-read sequencing that can span entire repeat blocks and capture their methylation patterns and RNA products in full.
What this means for patients and future therapies
The study does not overturn the central role of the shortened D4Z4 region on chromosome 4 in FSHD, but it shows that this region lives in a crowded neighborhood of similar sequences. Signals once assumed to prove activity at the disease locus may in fact come partly from its genomic cousins. For patients, this means that the most reliable diagnostic tools will be those that physically resolve the exact chromosome and repeat array involved, rather than relying solely on indirect readouts. For researchers and drug developers, it highlights the need for highly specific designs that avoid accidentally targeting harmless repeats elsewhere in the genome. In the telomere-to-telomere era, understanding and treating FSHD will depend on seeing the full repeat landscape, not just a single landmark.
Citation: Salsi, V., Losi, F., Pini, S. et al. Rethinking genomics of facioscapulohumeral muscular dystrophy in the telomere-to-telomere era: pitfalls in the hidden landscape of D4Z4 repeats. Eur J Hum Genet 34, 357–367 (2026). https://doi.org/10.1038/s41431-025-02000-x
Keywords: facioscapulohumeral muscular dystrophy, D4Z4 repeats, DUX4, telomere-to-telomere genome, genetic diagnostics