Clear Sky Science · en
Epigenetic context defines the transcriptional activity of canonical and noncanonical NF-κB signaling in pancreatic cancer
Why this study matters for patients
Pancreatic cancer is one of the deadliest cancers, in part because its cells are experts at reading and responding to danger signals in their surroundings. This study looks at two such signals, called TNFα and TWEAK, and shows how they switch on different “reading modes” in cancer cell DNA. Understanding these modes could open new paths to slow tumor growth, limit spread, and make treatments more effective.
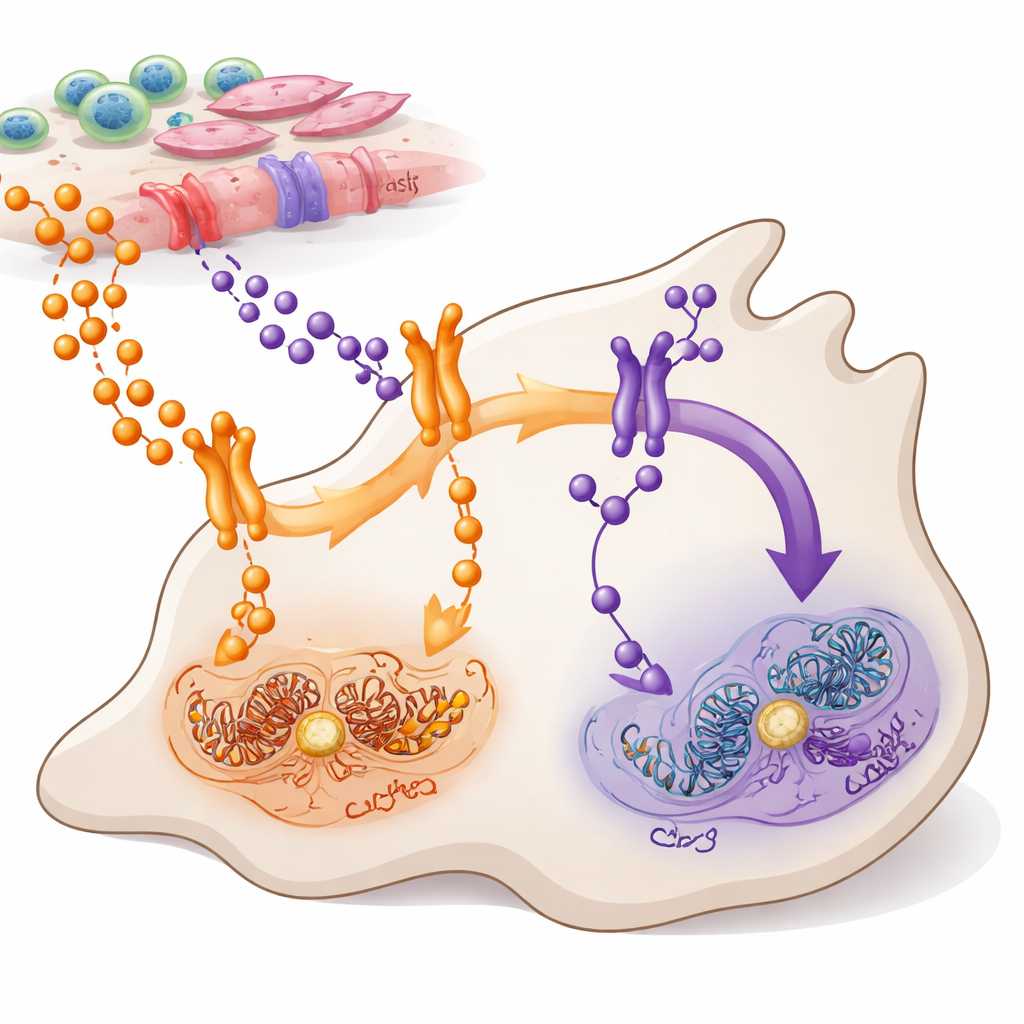
Two message routes into cancer cells
TNFα and TWEAK are small protein messengers released by cells in and around a tumor. Both talk to a major control system inside cells known as NF-κB, which helps decide whether cells grow, move, or die. The authors focused on pancreatic ductal adenocarcinoma, the most common and aggressive form of pancreatic cancer, where NF-κB is abnormally active in most tumors. They mapped which cells produce TNFα and TWEAK, which cells carry their receptors, and how these signals flow through two related NF-κB branches—often called the canonical (via RELA) and noncanonical (via RELB) routes.
Who sends the signals in the tumor neighborhood
By combining single-cell RNA sequencing from thousands of cells in patient tumors with advanced microscopy, the team found that TNFα mainly comes from immune cells, especially macrophages and certain T and B cells. TWEAK, in contrast, is produced by a broader cast: macrophages, fibroblasts, endothelial cells and stellate cells. Their receptors also differ. Tumor cells and nearby fibroblasts strongly express the main TNFα receptor and the key TWEAK receptor. Network analysis suggested that TWEAK-based signaling forms a wider and more complex communication web across tumor and support cells, while TNFα signaling is more focused within immune-rich areas.
Different depths of response inside tumor cells
In pancreatic cancer cell lines, the researchers treated cells with TNFα or TWEAK and measured which genes switched on over time. TNFα triggered a rapid and broad wave of gene activity linked to inflammation, cell movement, tissue remodeling, and survival. TWEAK induced a smaller, slower set of changes that largely overlapped with TNFα’s targets instead of launching a unique program. Both signals could promote cell migration and, under certain conditions, cell death, but TNFα had the stronger effect on movement. When the team examined patient tumor data from The Cancer Genome Atlas, they saw that tumors with high TNFα or high TWEAK levels shared many activated genes, confirming these patterns in real human cancers.
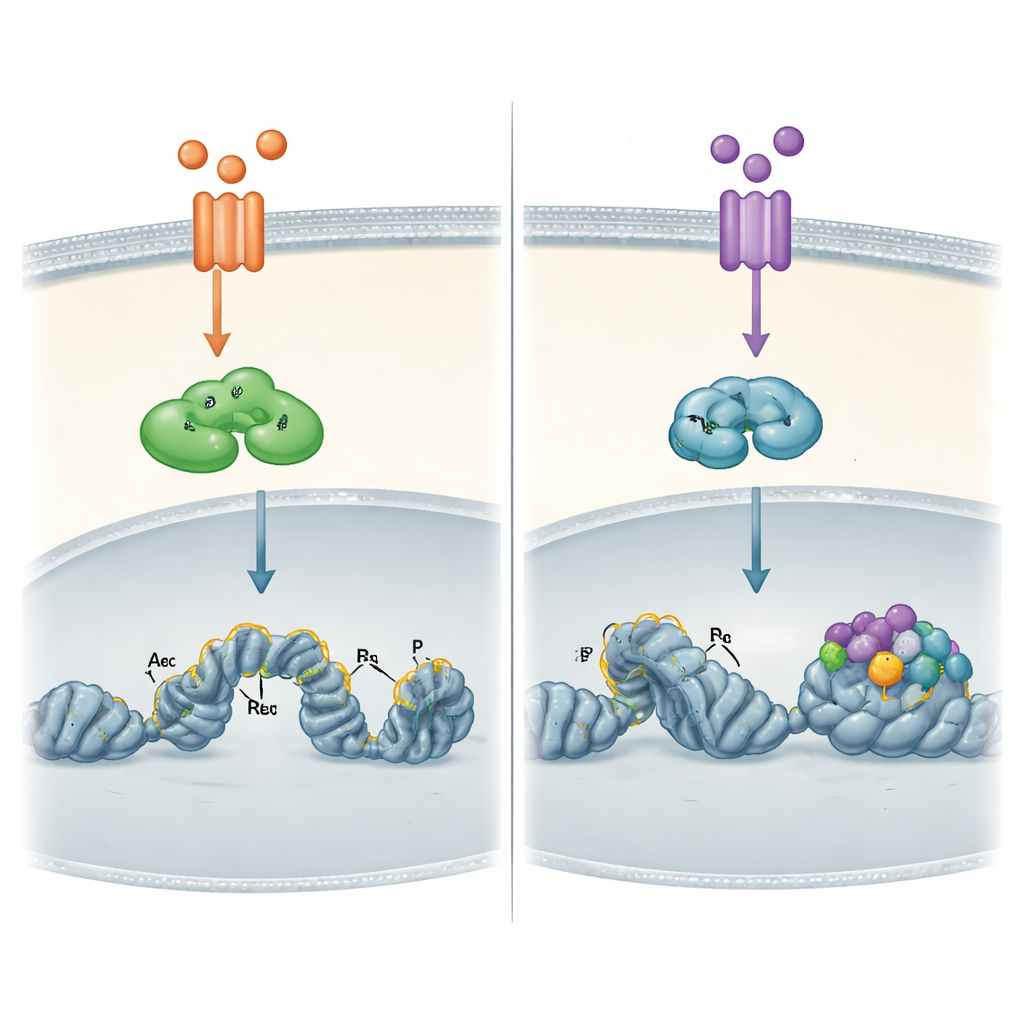
How the DNA landscape shapes each pathway
The most striking differences emerged when the authors looked directly at the cancer cells’ chromatin—the packed form of DNA and its associated proteins. Using genome-wide binding maps, they showed that TNFα-activated RELA can bind not only to already open DNA regions but also to more tightly packed regions and then help “open” them by adding chemical marks associated with active switches. RELB, turned on later by TWEAK, bound almost exclusively to sites that were already accessible and decorated with such active marks. These RELB sites were especially rich in docking motifs for another factor family, AP-1, suggesting that RELB depends on other proteins to prepare the ground before it can act.
What this means for future treatments
For a non-specialist, the key takeaway is that TNFα and TWEAK use the same NF-κB toolkit in very different ways. TNFα, through RELA, behaves like a master switch that can pry open closed sections of the cancer cell’s DNA and activate a wide range of genes involved in growth, survival, and spread. TWEAK, through RELB, is more like a specialist that can only operate where the DNA is already open and where co-factors such as AP-1 are present. This division of labor suggests that drugs targeting chromatin remodeling, AP-1, or specific NF-κB branches could selectively blunt harmful gene programs in pancreatic cancer while leaving other functions more intact.
Citation: Aggrey-Fynn, J.E., Busch, J., Saul, D. et al. Epigenetic context defines the transcriptional activity of canonical and noncanonical NF-κB signaling in pancreatic cancer. Cell Death Discov. 12, 152 (2026). https://doi.org/10.1038/s41420-026-03019-9
Keywords: pancreatic cancer, NF-kappaB signaling, tumor microenvironment, epigenetic regulation, TNF and TWEAK