Clear Sky Science · en
Concordance analysis of DNA and RNA profiling: The MD Anderson IMPACT2 study in precision oncology
Why this research matters for cancer patients
Cancer treatment is increasingly guided by the genetic makeup of each person’s tumor. Doctors already use DNA tests to find mutations that can be targeted with specific drugs, but these tests do not tell the whole story about how a tumor behaves. This study from MD Anderson Cancer Center asks a practical question: if we also look at RNA—the “working copies” of genes that show which genes are actually switched on or off—can we better understand a patient’s cancer and their chances of survival?
Two layers of information: DNA and RNA
Every cancer cell carries changes in its DNA, but DNA is mostly a static blueprint. RNA, in contrast, captures what the cell is actively doing at a given moment. In the IMPACT2 trial, more than 800 patients with advanced cancers had their tumors genetically profiled. For 253 of them, the researchers had both DNA and RNA data. They compared which genes were altered at the DNA level (such as mutations or extra copies) and which genes showed unusually high or low RNA levels, asking how often these two layers told the same story and whether that information related to how long patients lived.
When gene changes and activity line up
The team first looked for “concordant” events—cases where the same gene was abnormal in both its DNA and its RNA. Among the 253 patients, 50 had at least one such match, for a total of 58 events across 23 genes. Most of these involved extra or missing copies of a gene that also showed higher or lower RNA levels, and this alignment was strongest for known cancer drivers such as CDKN2A, AR, ESR1, KRAS, PIK3CA, AKT2, TP53, and CCND1. These findings support the idea that, for some key cancer genes, structural DNA changes do indeed translate into increased or decreased gene activity, reinforcing their importance as treatment targets.
Hidden relationships revealed by RNA signals
Beyond one-to-one matches, the researchers tested more than 12,000 gene pairs to see whether a DNA change in one gene was consistently linked with abnormal RNA levels in another. They found 123 significant pairs. Many of these clustered in a major growth and survival network known as the PI3K/AKT pathway, a frequent focus of targeted drugs. An especially notable pattern connected changes in the tumor-suppressor gene TP53 with overactive RNA signals from VEGFA, a gene that helps tumors build new blood vessels. This relationship could help explain why some patients whose tumors carry TP53 alterations respond to bevacizumab, a drug that blocks blood-vessel growth, and illustrates how RNA data can expose clinically relevant gene–gene interactions that DNA alone may miss.
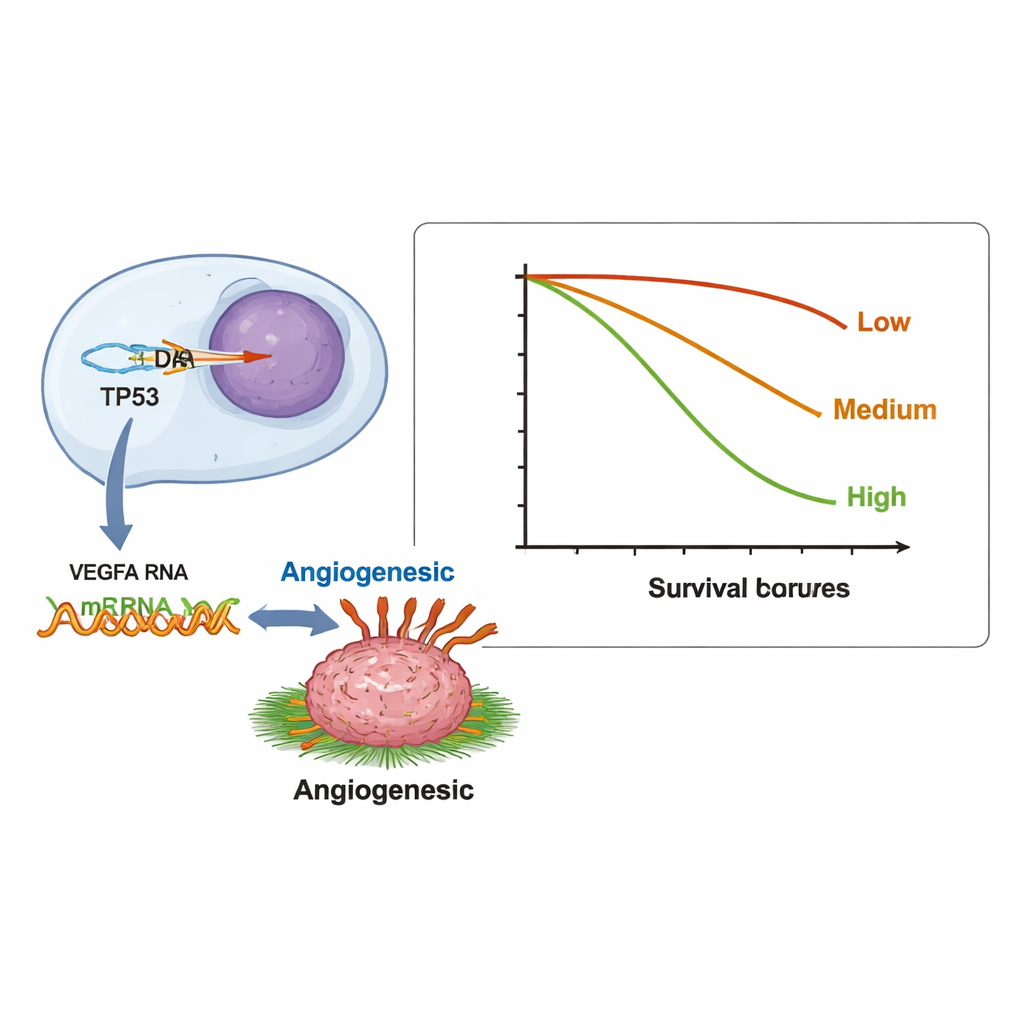
Gene activity load and patient survival
The investigators also asked whether the overall amount of abnormal gene activity in a tumor—what they call tumor transcriptional burden, or TTB—was linked to how long patients survived. They counted how many genes showed altered RNA levels in each patient and grouped them into low (0–2 genes), intermediate (3–5), and high (6 or more). Patients in the high group lived a median of 6.7 months, compared with 9.8 and 11.9 months in the lower groups. In other words, tumors with many misbehaving genes at the RNA level tended to be more aggressive. Tumors that lacked PD-L1, a marker often used to select patients for immunotherapy, also tended to have more genes with abnormal expression, hinting at a possible link between broad gene disruption and a tumor environment that resists immune attack.
What this means for future cancer care
This study shows that RNA profiling adds useful, complementary information to DNA testing in real-world patients with advanced cancers. In some genes, DNA changes and RNA activity line up and reinforce known treatment targets; in others, RNA patterns uncover new relationships that might guide drug choices, such as the tie between TP53 and VEGFA. Most importantly for patients, a heavy burden of abnormal gene activity was associated with shorter survival, suggesting that RNA-based measures could help doctors gauge how aggressive a tumor is and refine prognosis. While RNA testing is not yet routinely used to pick therapies, these findings support its promise as a next step in precision oncology, especially as larger studies and better analytical tools help bring transcriptomic profiling into everyday cancer care.
Citation: Schmidt, S.T., Baysal, M.A., Fu, S. et al. Concordance analysis of DNA and RNA profiling: The MD Anderson IMPACT2 study in precision oncology. Sig Transduct Target Ther 11, 68 (2026). https://doi.org/10.1038/s41392-026-02580-0
Keywords: precision oncology, RNA profiling, DNA sequencing, tumor transcriptional burden, cancer biomarkers