Clear Sky Science · en
The consensus molecular subtypes of esophageal squamous cell carcinoma
Why this research matters for patients
Esophageal squamous cell carcinoma is a deadly throat cancer with few tailored treatment options and generally poor survival. Doctors know that not all tumors behave the same, but until now there has been no clear, unified way to sort these cancers into biologically meaningful groups. This study pulls together many strands of genetic and imaging data to define four major types of this cancer and shows that a simple microscope slide image can often reveal which type a patient has. That opens the door to more precise, less trial‑and‑error treatment.
Putting order into a confusing cancer landscape
Over the past decade, different research teams proposed at least seven ways to divide this cancer based on DNA mutations, copy‑number changes, gene activity, or other molecular measurements. Each system used different data and methods, leading to overlapping but not identical subtypes and making it hard to use in everyday care. The authors first re‑analyzed a well‑characterized group of 152 Chinese patients for whom multiple layers of data were available, including whole‑genome sequencing, gene activity, DNA methylation, and microRNA profiles. Using a method called similarity network fusion, they combined all these data into a single picture and identified four multi‑omics subtypes. They then built a statistical “network of subtypes” that linked their new groups to 21 previously published ones, revealing strong connections across studies.
Four main biological personalities of the tumor
From this network, the team distilled four consensus molecular subtypes, named ECMS1 through ECMS4, each with distinct behavior. ECMS1, the metabolic type (MET), shows high activity in pathways that process drugs and nutrients and frequent activation of the NFE2L2 gene, which can drive resistance to chemotherapy and radiotherapy. ECMS2, the classical type (CLS), is dominated by strong cell‑cycle and growth‑signal activity, and often carries extra copies of the ERBB2 (HER2) gene. ECMS3, the immune type (IM), has lower overall mutation burden but strong immune‑system activation, including high levels of the PD‑1 checkpoint pathway. ECMS4, the mesenchymal type (MES), shows signs of tissue remodeling, blood vessel growth, and cells becoming more mobile and invasive, features linked to worse outcomes.
Linking subtypes to prognosis and treatment choices
When the researchers compared these four groups with patients’ clinical data, clear patterns emerged. Patients with the mesenchymal ECMS4 tumors had the highest risk of recurrence and the poorest overall and disease‑free survival. The immune ECMS3 group, despite its active immune environment, responded poorly to standard chemoradiotherapy but showed a higher rate of benefit from an anti‑PD‑1 immunotherapy drug. The classical ECMS2 tumors appeared more sensitive to common chemotherapy drugs such as cisplatin and taxanes, and their frequent HER2 amplification suggests they may benefit from HER2‑targeted medicines. The metabolic ECMS1 tumors, enriched for NFE2L2 activity and certain growth‑factor receptors, may be better treated with drugs targeting those metabolic and signaling pathways. In short, the four subtypes suggest different weak points and different likely responses to therapy.
Teaching a computer to see molecular clues in routine slides
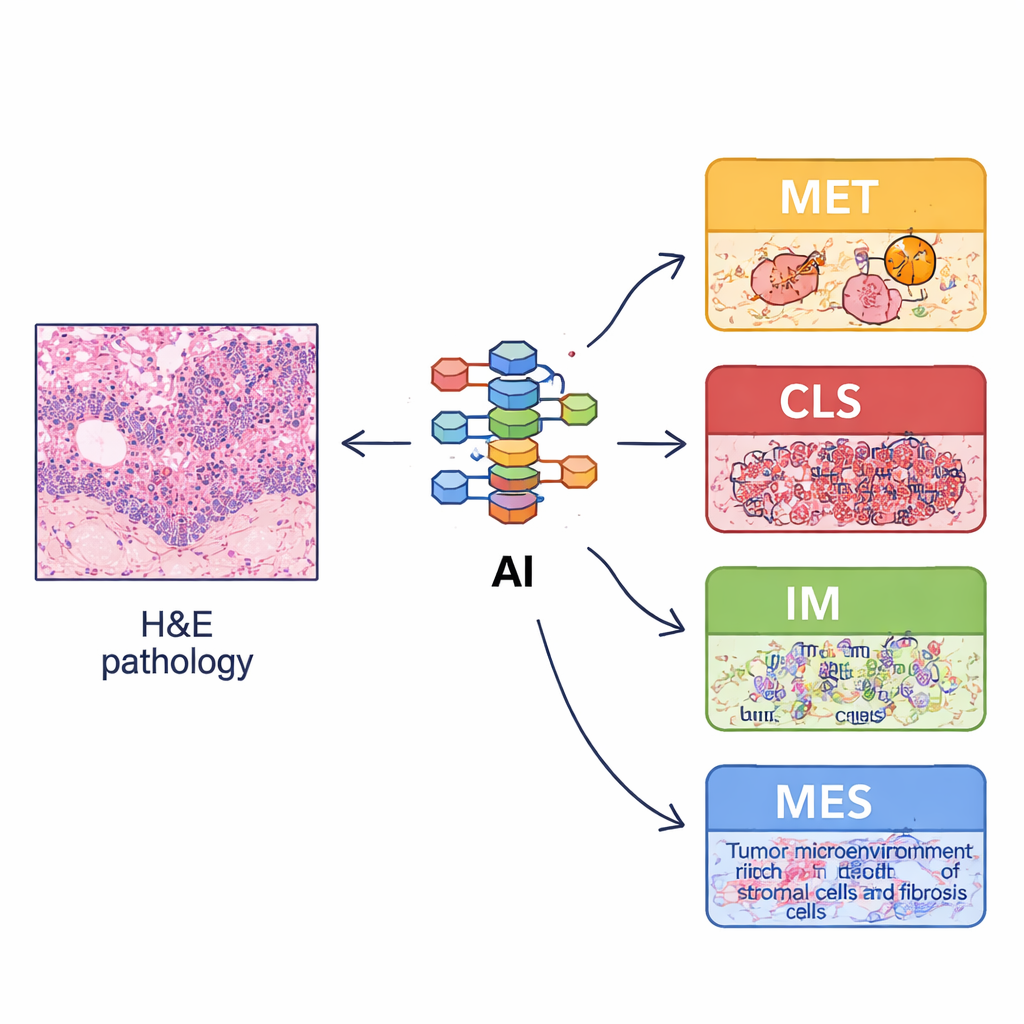
Because full molecular profiling is expensive and not available in many hospitals, the team asked whether ordinary hematoxylin‑and‑eosin (H&E) pathology slides could stand in as a proxy. They trained a deep‑learning system on tens of thousands of annotated image snippets to recognize basic tissue elements such as tumor cells, stroma, lymphocytes, glands, and muscle. From whole‑slide images, they then computed “spatial organization features” – how much of each tissue type is present, how they are arranged, and how mixed or separated they are. These image‑derived patterns mirrored the biology of the four subtypes: for example, the immune ECMS3 type showed more lymphocytes and normal structures, while the mesenchymal ECMS4 type showed more supporting stroma. Using these features, they built an image‑based classifier, called imECMS, that could assign slides to MET, CLS, IM, or MES with good accuracy in several independent patient groups.
What this means for future care
For non‑specialists, the takeaway is that this study transforms a seemingly uniform cancer into four biologically distinct diseases, each with its own likely course and best treatment strategies. Even more importantly, it shows that much of this information can be extracted automatically from standard microscope images that every patient already has, without requiring costly sequencing. With further validation in larger and more diverse populations, the ECMS and imECMS systems could help doctors match patients with esophageal squamous cell carcinoma to the therapies they are most likely to benefit from, while sparing them from treatments unlikely to work.
Citation: Cui, H., Zhu, Z., Xu, E. et al. The consensus molecular subtypes of esophageal squamous cell carcinoma. Sig Transduct Target Ther 11, 65 (2026). https://doi.org/10.1038/s41392-026-02577-9
Keywords: esophageal cancer, tumor subtypes, precision oncology, deep learning pathology, immunotherapy