Clear Sky Science · en
Common gene mutations in 103 authenticated colorectal cancer cell lines
Why these cancer cell lines matter
Colorectal cancer is one of the most common cancers worldwide, and much of what we know about how it behaves and how to treat it comes from studies in laboratory-grown cancer cell lines. But these models can drift and change over time. This study carefully checked and cataloged 103 widely used colorectal cancer cell lines, mapping out key gene mutations that drive tumor growth. The result is a detailed "field guide" that helps researchers choose the right model for the right question, ultimately aiming to make lab discoveries more likely to benefit future patients.
A catalog of cancer models
The researchers assembled 103 colorectal cancer cell lines that together mirror the main genetic subtypes seen in patients. Most came from primary colon or rectal tumors, with the rest from metastases. The team first verified that each cell line was truly what it claimed to be, using fingerprint-like DNA markers called short tandem repeats. They then used deep targeted sequencing, reading selected stretches of DNA on average 575 times, to examine 20 genes known to be important in colorectal cancer. This high depth allowed them to detect even relatively rare mutations within a cell population and to estimate how common each mutation was among the cells in a given line.
Different paths to genetic chaos
Colorectal cancers can take different genetic routes as they develop. About four out of five tumors show chromosomal instability, with large chunks of DNA gained or lost. Others are “hypermutated,” piling up many small DNA changes. This study captured both patterns. Sixteen cell lines showed microsatellite instability, a form of hypermutation caused by faulty DNA mismatch repair, leading to frequent small insertions and deletions. Another five had defects in the DNA proofreading gene POLE, generating a heavy load of single-letter DNA changes. The remaining, non-hypermutated lines had fewer mutations but more frequent large-scale DNA copy number changes. These distinct signatures closely resembled those seen in real patient tumors.
Key driver genes and dangerous combinations
Across the 20 genes surveyed, the mutation patterns matched large clinical studies of colorectal cancer. Tumor-suppressor genes such as APC, TP53, and SMAD4 were commonly hit by truncating mutations that break the protein, while classic cancer-promoting genes like KRAS and BRAF more often carried single amino-acid changes that switch signaling pathways on. Non-hypermutated lines tended to carry a higher share of clearly harmful, or pathogenic, mutations that were present in most or all cells in the culture. In contrast, hypermutated lines carried many extra “passenger” changes and more subclonal mutations that appear only in a fraction of cells, reflecting ongoing evolution in the dish.
Hidden patterns in how genes interact
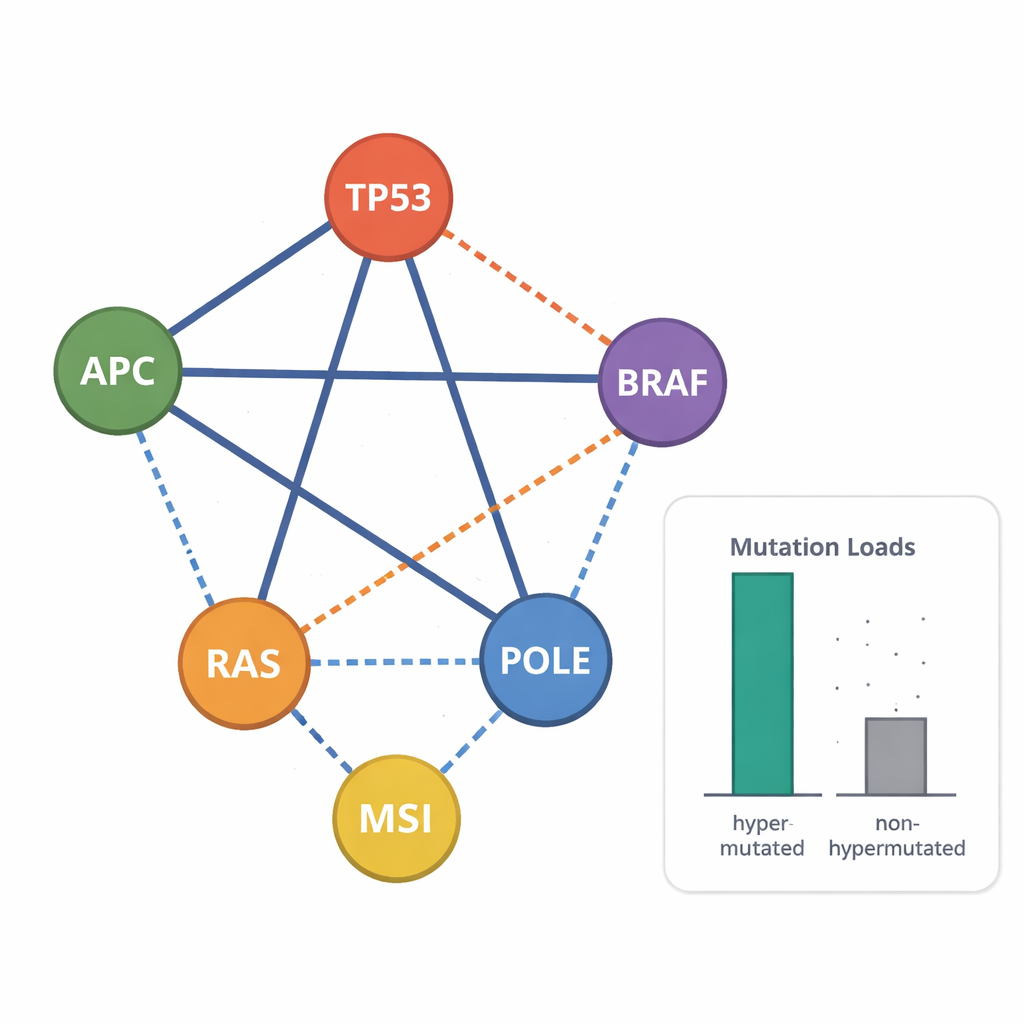
By looking at which mutations tend to appear together or avoid each other, the scientists could spot genetic combinations linked to particularly aggressive disease. For example, the common hotspot mutation BRAF p.V600 rarely co-occurred with KRAS or NRAS mutations, but in a handful of lines it did appear alongside truncating APC mutations, echoing a poor-prognosis subtype seen in patients. Many cell lines had triple hits in APC, TP53, and RAS genes, another marker of high-risk tumors. The study also uncovered a distinctive “double-hit” pattern in APC: two truncating mutations placed so that at least one β-catenin–binding region remains, consistent with a “just-right” level of WNT pathway activation that favors tumor growth. Copy number analysis showed frequent amplifications of growth-related genes like MYC and EGFR, and loss of entire copies of tumor-suppressor genes, especially in non-hypermutated lines.
What this means for future research
For scientists designing experiments, the message is that not all colorectal cancer cell lines are created equal. Hypermutated models are genetically busy and may contain many low-level changes that blur the impact of any single mutation. Non-hypermutated lines, by contrast, tend to carry fewer but stronger and more uniform driver alterations. By providing a carefully validated map of which genes are altered, how they are altered, and how common each mutation is within each culture, this work equips researchers to pick the most appropriate models for testing drugs, probing biology, and developing new targeted or immunotherapy approaches for colorectal cancer.
Citation: Kranjec, C., Eilertsen, I.A., Nunes, L. et al. Common gene mutations in 103 authenticated colorectal cancer cell lines. Oncogenesis 15, 8 (2026). https://doi.org/10.1038/s41389-026-00599-0
Keywords: colorectal cancer, cancer cell lines, gene mutations, tumor subtypes, precision oncology