Clear Sky Science · en
Strategies to control cellular spatial organization in microphysiological systems
Building Tiny Tissues in the Lab
Our organs work so well because their cells are not randomly scattered—they are carefully arranged in space. This paper explores how scientists are learning to recreate that intricate order inside “organs-on-chips” and other miniature lab-grown tissues. By guiding where different cells sit and how they interact, researchers can build more realistic models of the heart, brain, gut, blood vessels, and even human–microbe ecosystems. These advanced systems promise safer drug testing, fewer animal experiments, and better tools to study diseases and develop personalized treatments.
Why Cell Position Matters
Inside the body, every cell lives in a specific neighborhood with its own mix of support structures, chemical signals, and physical forces. A liver cell near a blood vessel feels very different cues than a cell buried deep in the organ. These differences in location help determine how cells grow, what they turn into, and how they respond to injury or medicine. When scientists grow cells on flat plastic dishes, that spatial “conversation” is mostly lost, and cells often behave in unnatural ways. The paper argues that recreating realistic spatial organization is not an optional upgrade but a basic requirement if lab-grown tissues are to mimic real organs.
Two Main Ways to Arrange Cells
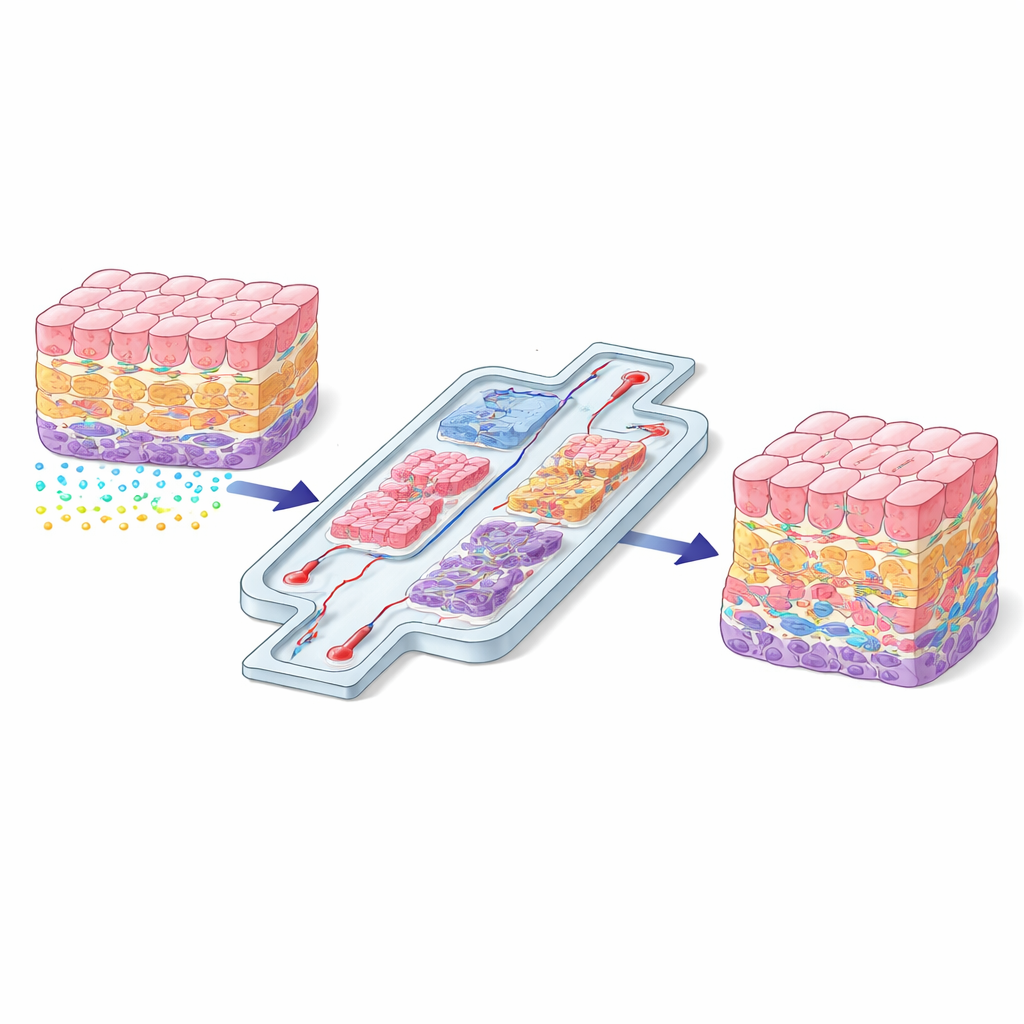
The authors group current strategies into two broad families: direct and indirect control. Direct methods physically place cells or compartments exactly where they are needed. Examples include 3D bioprinting, which lays down layers of cells and soft gels like a biological 3D printer; microfluidic chips that carve tissues into connected chambers and channels; and physical trapping methods that nudge cells into position using light, sound, magnets, or electric fields. These approaches shine when precise geometry is crucial—such as recreating a blood–brain barrier, a layered blood vessel wall, or the flow path from gut to liver.
Letting Cells Read Their Surroundings
Indirect methods instead reshape the environment and let cells do the rest. Here, scientists tune the “landscape” that cells sense: the makeup of the surrounding gel, the stiffness and texture of surfaces, and gradients of dissolved substances like growth factors or drugs. Carefully arranged support molecules can make certain cell types settle in specific regions. Smart hydrogels that soften or release signals when cells remodel them encourage self-organized structures such as blood vessel networks or gut-like crypts and villi. Microfluidic devices can generate stable chemical gradients that cause stem cells to adopt different identities along a channel, or cancer cells and immune cells to migrate into distinct zones, revealing how diseases spread.
From Single-Cell Systems to Whole-Body Models
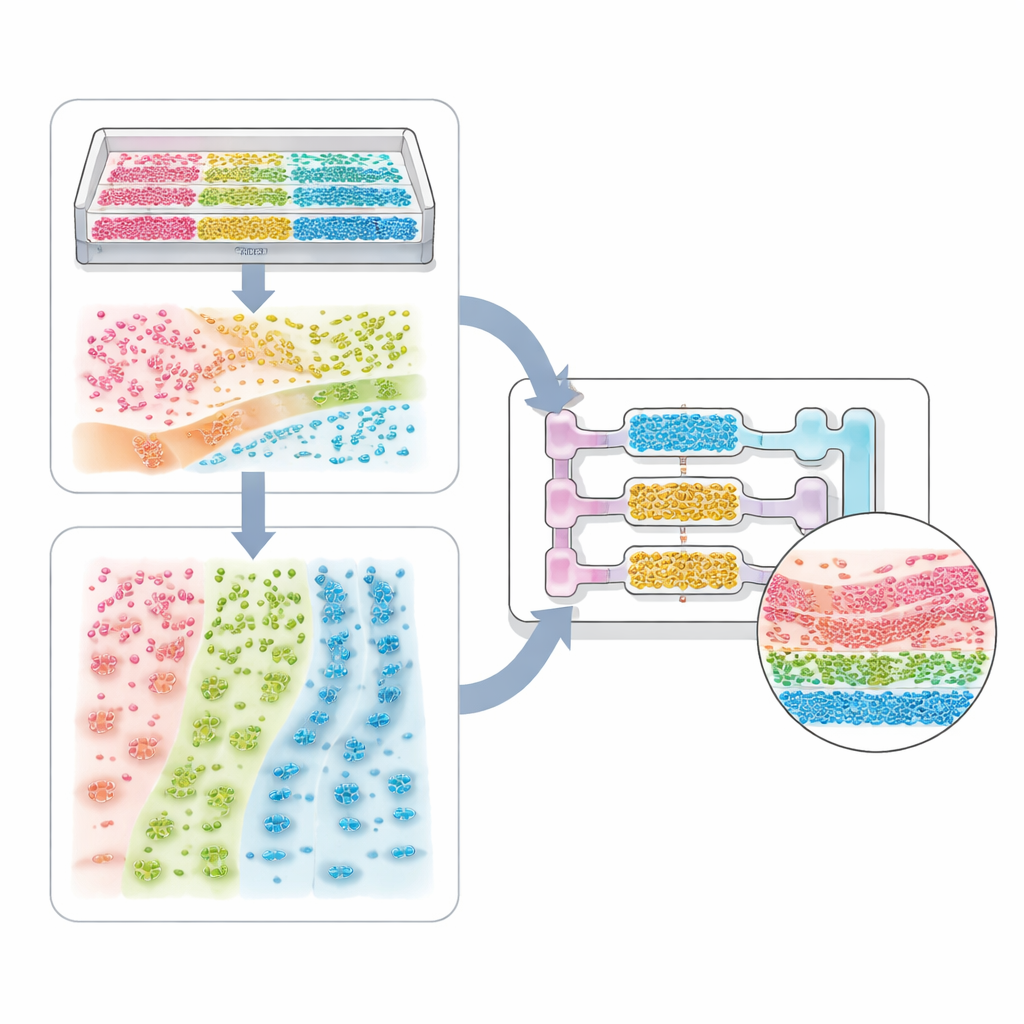
The review walks through a spectrum of applications. At the simplest end are single cell-type systems, where patterned grooves or stiffness gradients can make heart cells beat in unison or nerve cells extend aligned fibers. More complex tissue models mix several cell types—such as tumor cells, support cells, and blood vessel cells—to form spheroids and organoids with internal cores and shells. Microchannel chips and bioprinting then add boundaries and flow, allowing the recreation of lung air sacs, kidney filters, liver zones, and multi-organ circuits that track how a drug is absorbed, transformed, and cleared. The same ideas extend to host–microbe systems, where the spatial arrangement of bacteria along the gut wall, or across a mucus layer, can determine whether they protect us or cause disease.
Checking That Patterns Are Real
Because these systems are getting more intricate, scientists need reliable ways to confirm that cells end up where they should and behave as intended. The paper highlights imaging methods that can watch living cells move and change over time, as well as advanced staining and sequencing techniques that map which genes, proteins, and metabolites appear at each location. Sensors embedded in chips can track oxygen, nutrients, acids, and mechanical forces, tying local conditions to cell responses. Together, these tools help researchers verify that a design is not just visually convincing but functionally faithful to real tissue.
Where This Work Is Heading
The authors conclude that the most powerful systems will blend direct and indirect strategies: using chips and bioprinters to set the overall layout, then layering in tunable gels, textures, and gradients that let tissues mature and remodel over time. They also note practical hurdles—such as scaling up fabrication, reducing variability between labs, and addressing ethical questions around complex human tissue models. Still, the message is clear: mastering spatial organization is key to turning microphysiological systems into trustworthy stand-ins for human organs, opening new doors for studying development, testing therapies, and tailoring medicine to individual patients.
Citation: Truong, H.D., Ge, Z., Chng, E. et al. Strategies to control cellular spatial organization in microphysiological systems. Microsyst Nanoeng 12, 85 (2026). https://doi.org/10.1038/s41378-025-01141-9
Keywords: organ-on-a-chip, cell spatial organization, 3D bioprinting, microfluidic tissue models, microphysiological systems