Clear Sky Science · en
One-pot CRISPR-based point of care platform for rapid, specific and sensitive detection of HPV 16 without pre-amplification
Why spotting one virus type really matters
Cervical cancer is often linked to infection with human papillomaviruses (HPV), but not all HPV types carry the same risk. Doctors therefore need tests that can reliably tell these very similar viruses apart, especially the high‑risk type called HPV16. The challenge is that today’s genetic tests can sometimes confuse close relatives, leading to false alarms or missed cases. This study introduces a new testing method that sharply improves both accuracy and practicality, aiming to bring precise HPV subtyping to clinics and point‑of‑care settings.
A smart molecular lock and key
The researchers built a one‑tube test they call CASTSA that combines two powerful ideas: the precision of CRISPR, a gene‑targeting system adapted from bacteria, and the signal‑boosting ability of PCR, the standard method for copying DNA. In CASTSA, a CRISPR protein named Cas12a is guided to the HPV16 genetic sequence by a short RNA “key.” When it finds the exact match plus a nearby short pattern it requires, Cas12a cuts the viral DNA at a predictable spot. This cut creates a distinctive loose end on one strand of DNA that serves as a unique starting point for amplification. Other HPV types, even with very similar sequences, are either not cut or do not generate the right starting point, so they do not get amplified.
Turning a tiny cut into a strong signal
To convert that CRISPR cut into a detectable signal, the team designed a special DNA primer called a terminal‑specific primer. This primer only attaches properly to the strand produced by Cas12a’s cut, then folds back on itself like a hairpin and helps build a stable template for PCR. Additional “universal” primer sequences then latch on and drive a conventional real‑time PCR reaction. Crucially, amplification only happens if Cas12a has first done its job and if the primer folds correctly, creating two layers of checking. This architecture sharply reduces the chance that partially matching, non‑target DNA will be accidentally amplified, addressing a common weakness of earlier CRISPR‑based tests that relied on a separate pre‑amplification step.
All in one pot, with fewer errors
A key practical advance is that CRISPR cutting and PCR amplification occur together in the same sealed tube, avoiding the need to open tubes between steps—a major source of contamination in many lab workflows. The authors carefully tuned reaction conditions, especially magnesium levels, so that both Cas12a and the DNA‑copying enzyme work well without encouraging spurious reactions. They showed that their one‑pot setup can faithfully distinguish HPV16 from several other high‑risk types, including HPV18, 33, 45, and 52, even though these viruses share highly conserved genetic regions. When compared to standard real‑time PCR alone, or to CRISPR tests that first amplify DNA and then perform CRISPR detection, CASTSA produced far fewer misleading signals from off‑target HPV types.
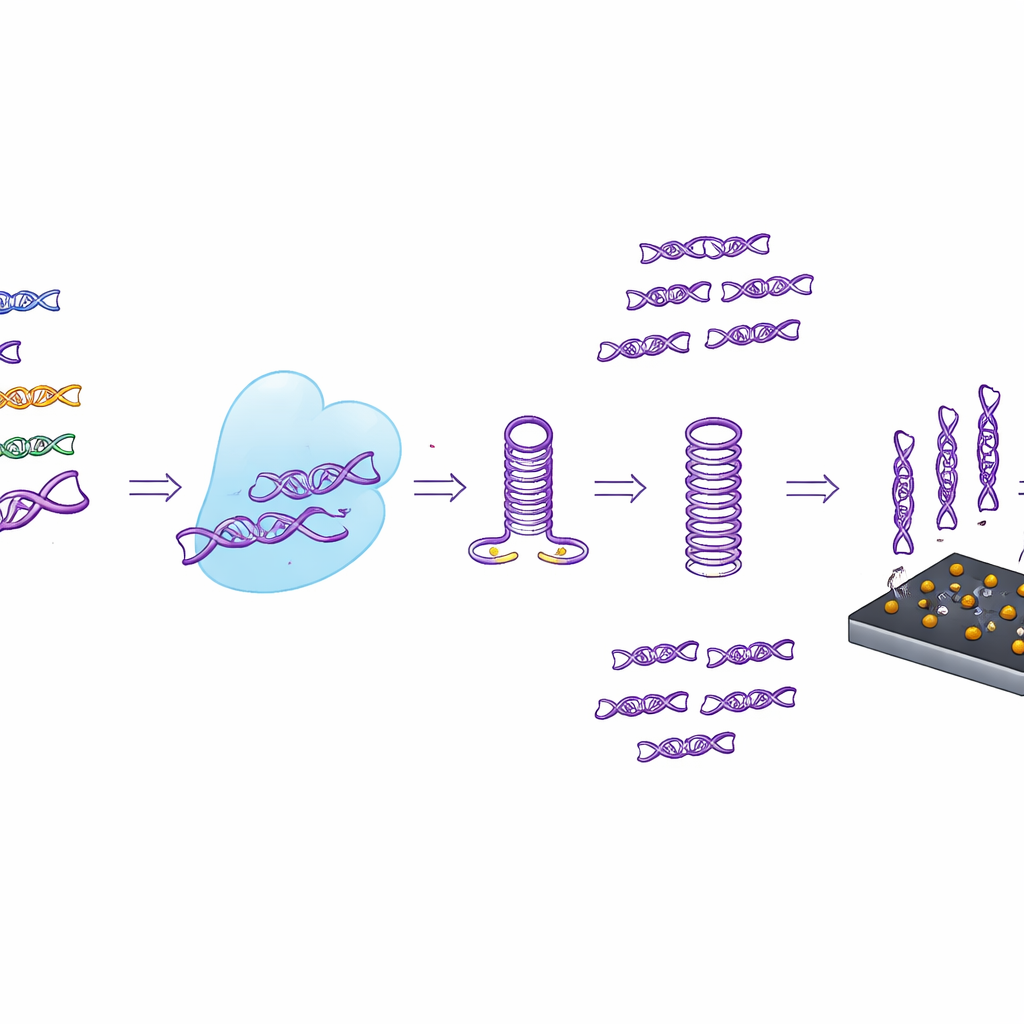
From test tube to handheld sensor
Instead of relying only on fluorescent light readouts, the researchers also coupled CASTSA to a compact electrochemical sensor made from laser‑induced graphene. This porous, highly conductive carbon surface is decorated with gold nanoparticles that carry capture strands designed to bind only the CASTSA amplification products from HPV16. When these products stick, the electrical current through the sensor shifts in a measurable way. By using a four‑electrode array—three measurement spots and one built‑in background control—the device can subtract noise from non‑specific sticking. In trials, this integrated system detected as few as 18 copies of the HPV16 target per reaction, outperforming the sensitivity of fluorescence alone and maintaining excellent repeatability.
How this could change HPV testing
To see how the method works in real life, the team tested 20 clinical samples known to contain HPV16 and 10 samples from healthy donors. CASTSA, both in its fluorescence form and when paired with the graphene‑based sensor, agreed completely with standard hospital PCR tests. At the same time, it showed superior ability to ignore other high‑risk HPV types that commonly cause confusion. For patients, this could mean fewer ambiguous results and more confident decisions about follow‑up and treatment. More broadly, the CASTSA strategy—CRISPR recognition first, amplification second, all in one sealed tube—offers a blueprint for highly specific, low‑contamination genetic tests that could be adapted to many pathogens and mutations, especially in settings that demand quick, reliable answers.
Citation: Chen, Y., Chen, Y., Zhang, C. et al. One-pot CRISPR-based point of care platform for rapid, specific and sensitive detection of HPV 16 without pre-amplification. Microsyst Nanoeng 12, 81 (2026). https://doi.org/10.1038/s41378-025-01130-y
Keywords: HPV16 detection, CRISPR diagnostics, PCR-based testing, electrochemical biosensor, point-of-care assay