Clear Sky Science · en
Structural insights into disease-associated mutations in the microRNA processing machinery
How Tiny RNA Scissors Shape Health and Disease
Inside every cell, a hidden editing system trims genetic messages before they are read. This system relies on tiny RNA snippets called microRNAs, which act like fine-tuning knobs for thousands of genes at once. When the molecular "machines" that make microRNAs work properly, cells grow, divide and specialize in a controlled way. But when these machines are damaged by mutations, the balance of gene activity can tip toward cancer, blood disorders or problems in brain development. This article explains how understanding the 3D shapes of these machines helps scientists pinpoint what goes wrong and how it might be fixed.
The Cell’s Gene-Dimming Toolkit
MicroRNAs are short pieces of RNA, about 22 building blocks long, that do not code for proteins themselves. Instead, they latch onto longer messenger RNAs and either mark them for destruction or block them from being translated, effectively turning genes down rather than fully off. To make these microRNAs, cells pass longer RNA precursors through a carefully staged assembly line. First, the enzyme DROSHA trims bulky primary transcripts in the cell nucleus into shorter hairpin-shaped pieces. These pieces then move to the cytoplasm, where another enzyme, DICER, measures and cuts them into microRNA-sized duplexes. Finally, one strand of each duplex is loaded into a protein called Argonaute 2 (AGO2), which uses the microRNA as a guide to find matching messages and silence them. 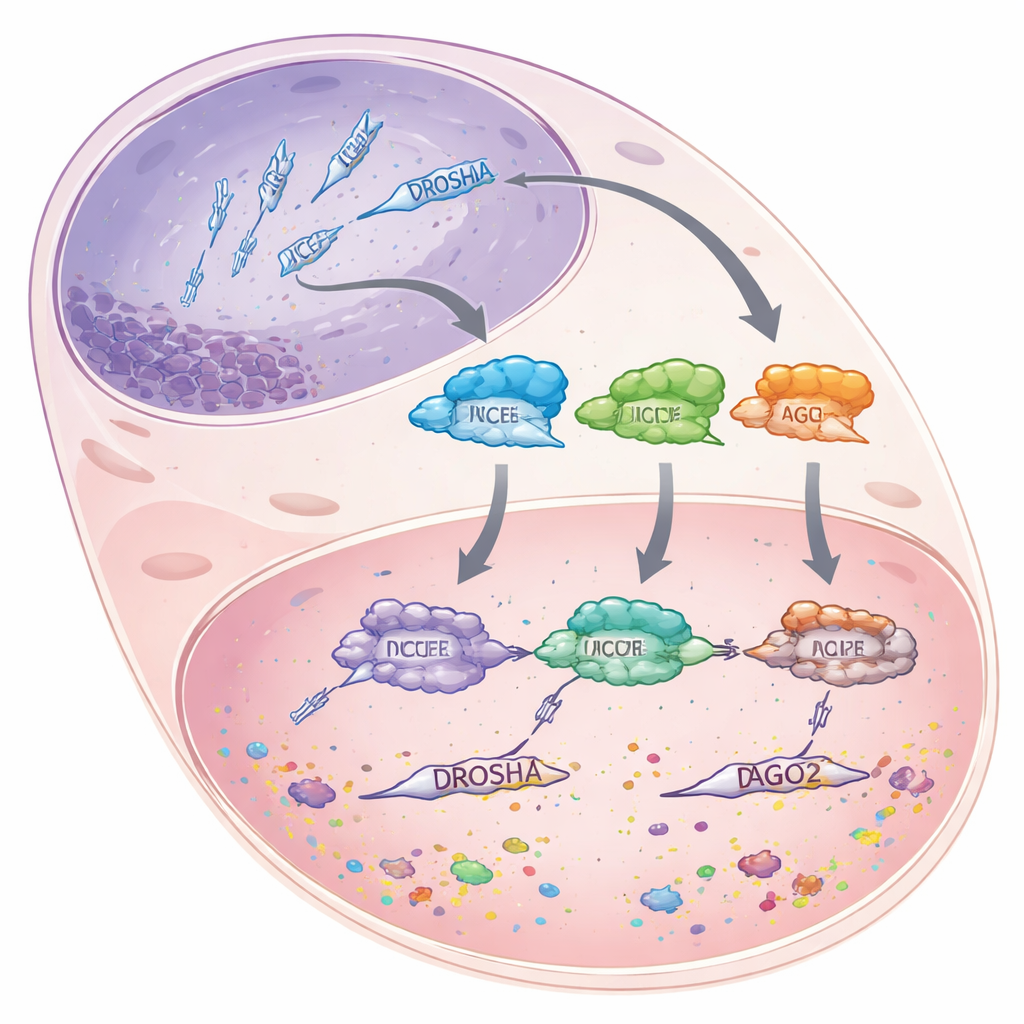
Delicate Shapes with Precision Jobs
DROSHA, DICER and AGO2 are not simple scissors; each is built from several interacting parts that must move in a choreographed way. DROSHA works together with a partner protein, DGCR8, to grip the ends of a primary microRNA hairpin and judge exactly where to cut. DICER uses a kind of built-in ruler formed by its domains to set a fixed distance between where it holds the hairpin end and where its catalytic center makes a cut. AGO2 wraps around a single microRNA strand so that its “seed” region is perfectly exposed for recognizing target messages, while metal ions and key amino acids in its core line up to slice those targets when pairing is strong enough. Across all three proteins, positively charged grooves and flexible linkers guide RNA strands into place while metal-binding pockets and catalytic residues carry out the chemical snips.
When the Scissors Are Bent or Blunt
Genetic studies in patients with cancer, blood disease and neurodevelopmental syndromes have uncovered clusters of mutations in these microRNA machines. Many of the most damaging changes lie directly in catalytic centers or in tight RNA-gripping surfaces. In DROSHA, for example, substitutions like E1147K and D1219G sit at the heart of the cutting site and disrupt the ability to hold essential metal ions, leaving the enzyme unable to process primary microRNAs. Other DROSHA mutations, such as truncations that remove the cutting region or changes that destabilize its support helices, weaken the whole complex and cause global drops in microRNA levels in Wilms tumor, bone marrow disorders and nervous system disease. In DICER, recurring mutations cluster in one of its two cutting domains, skewing production so that one arm of the microRNA duplex is lost, which shifts the balance between growth-promoting and growth-suppressing signals. 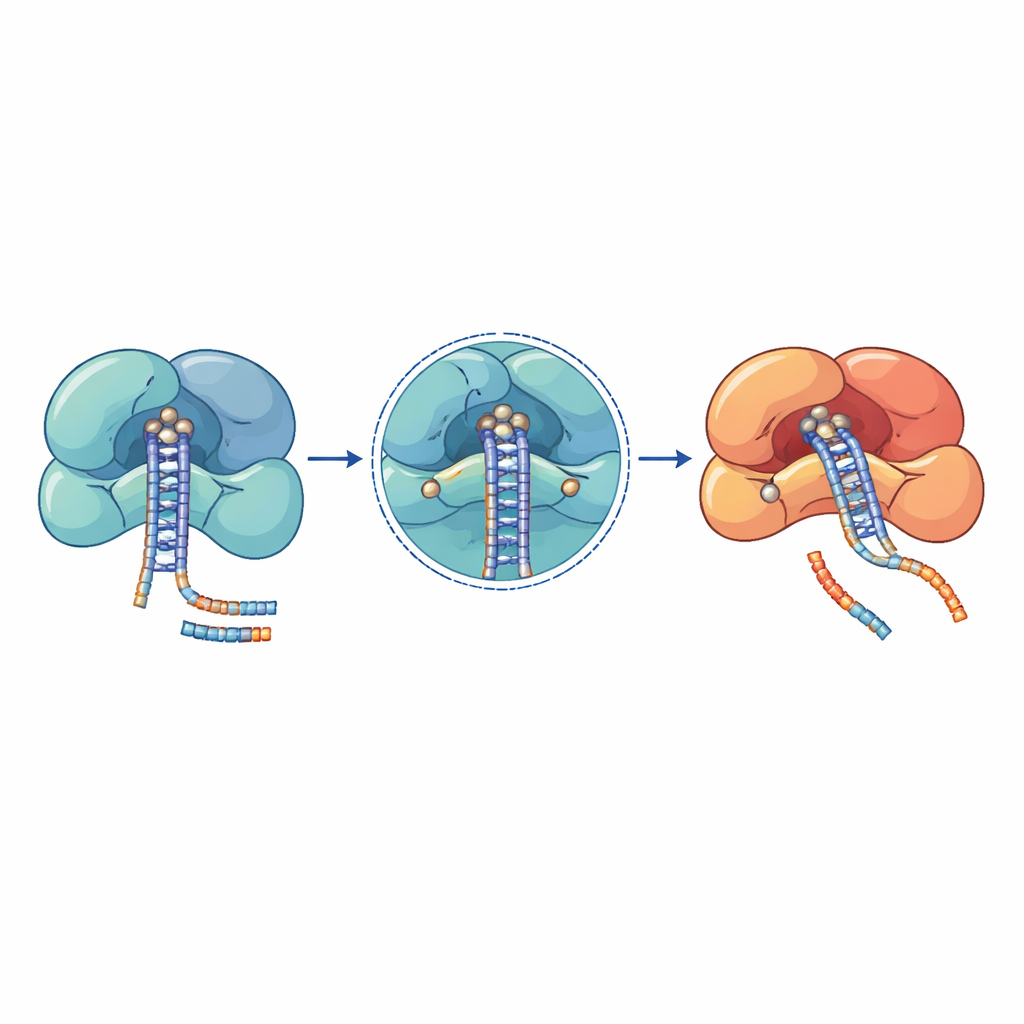
Misaligned Guides in the Brain and Beyond
AGO2 mutations tell a different but related story. Direct hits to its catalytic core are rare, likely because they would be lethal early in development. Instead, disease-linked variants tend to appear in flexible linker regions and at domain junctions that hold AGO2’s shape together. In Lessel–Kreienkamp syndrome, a neurodevelopmental condition, mutations in these linkers and surrounding surfaces subtly twist how the PAZ, MID and PIWI parts of AGO2 pack and move. This can misalign the guide microRNA or weaken its grip on target RNAs, blurring the precision of gene silencing during brain development. Similar structural changes anywhere along the microRNA pathway—whether in DROSHA, DICER or AGO2—can ripple outward, altering entire networks of genes involved in cell growth, blood formation or neuronal wiring.
Turning Structural Clues into Future Treatments
By overlaying disease mutations onto high-resolution 3D maps of these proteins, researchers can categorize exactly how each change causes trouble—whether by dulling the cutting edge, loosening critical contacts with partner proteins or bending key helices out of alignment. These structural insights open the door to targeted interventions: small molecules that stabilize weakened interfaces, designed RNAs that steer partially functional enzymes back onto the right substrates, or allosteric compounds that nudge mutant proteins into more active shapes. For a lay observer, the take-home message is that tiny structural tweaks in the cell’s RNA scissors can have outsized effects on health, but they also provide precise handles for designing next-generation, mutation-aware therapies.
Citation: Lee, H., Lee, J. & Roh, SH. Structural insights into disease-associated mutations in the microRNA processing machinery. Exp Mol Med 58, 639–649 (2026). https://doi.org/10.1038/s12276-026-01669-4
Keywords: microRNA, RNA interference, DROSHA DICER AGO2, cancer genetics, neurodevelopmental disorders